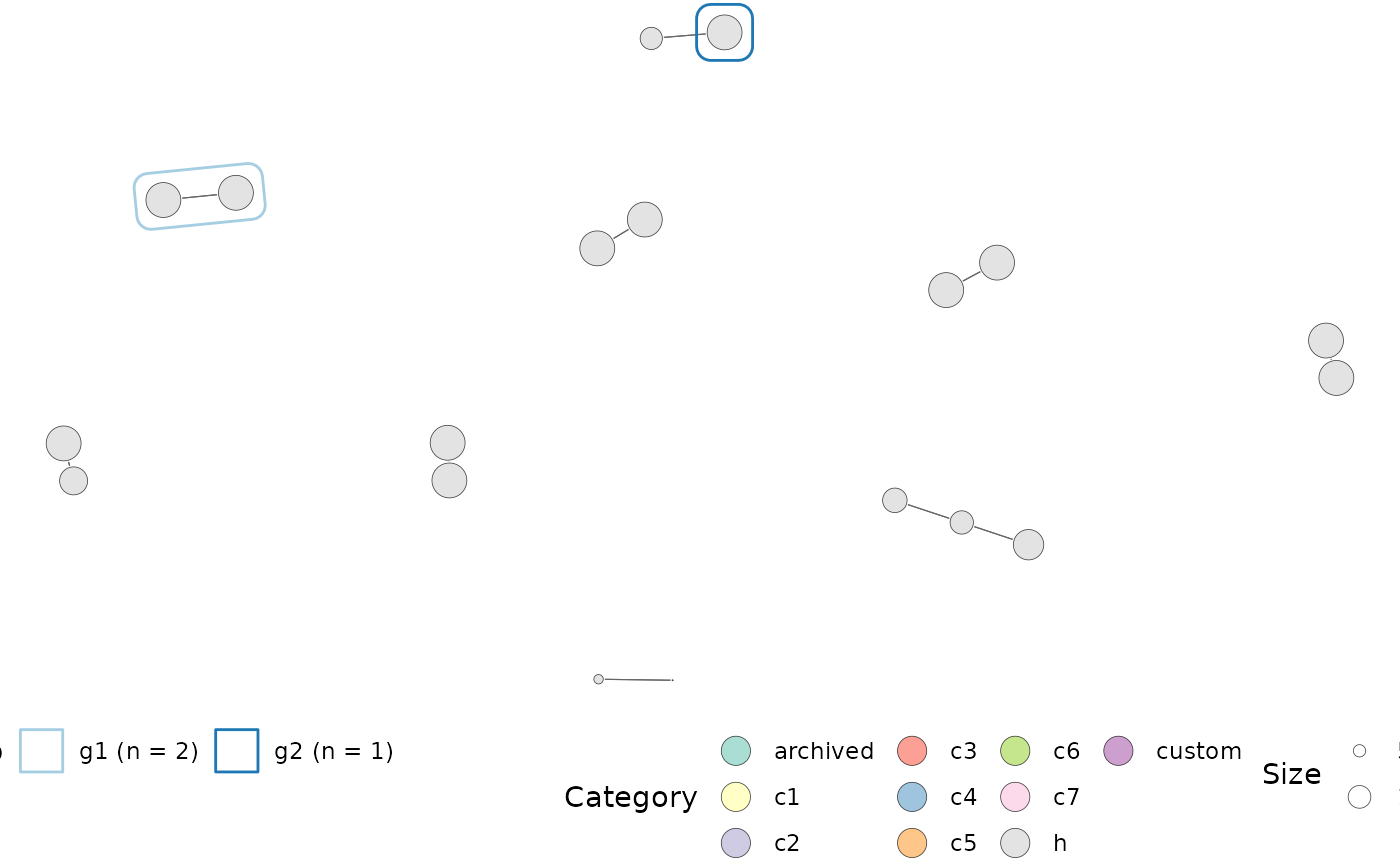
Plots a network of gene set overlap with overlap computed using the
computeMsigOverlap() and a graph created using computeMsigNetwork().
plotMsigNetwork(
ig,
markGroups = NULL,
genesetStat = NULL,
nodeSF = 1,
edgeSF = 1,
lytFunc = "graphopt",
lytParams = list(),
rmUnmarkedGroups = FALSE,
maxGrp = 12
)Arguments
- ig
an igraph object, containing a network of gene set overlaps computed using
computeMsigNetwork().- markGroups
a named list, of character vectors. Each element of the list represent a group and contains a character vector with node names. Up to 12 groups can be visualised in the plot.
- genesetStat
a named numeric, statistic to project onto the nodes. These could be p-values, log fold-changes or gene set score from a singscore-based analysis.
- nodeSF
a numeric, indicating the scaling factor to apply to node sizes.
- edgeSF
a numeric, indicating the scaling factor to apply to edge widths.
- lytFunc
a character, specifying the layout to use (see
ggraph::create_layout()).- lytParams
a named list, containing additional parameters needed for the layout (see
ggraph::create_layout()).- rmUnmarkedGroups
a logical, indicating whether unmarked groups should be removed from the network (TRUE) or retained (FALSE - default).
- maxGrp
a numeric, specifying the maximum number of groups to plot.
Value
a ggplot2 object
Examples
data(hgsc)
ovlap <- computeMsigOverlap(hgsc, thresh = 0.15)
ig <- computeMsigNetwork(ovlap, hgsc)
groups <- list(
'g1' = c("HALLMARK_HYPOXIA", "HALLMARK_GLYCOLYSIS"),
'g2' = c("HALLMARK_INTERFERON_GAMMA_RESPONSE")
)
plotMsigNetwork(ig, markGroups = groups)